Jason Shepherd

Professor of Neurobiology, Adjunct Associate Professor of Biochemistry, and Adjunct
Associate Professor of
Ophthalmology and Visual Sciences
Jon M. Huntsman Presidential Endowed Chair
Synaptic Plasticity, Memory and Neurological Disorders, Memory Formation
Education
B.Sc. 2002, University of Otago; Ph.D. 2007, Johns Hopkins University; Postdoctoral Fellow 2007-2012, Massachusetts Institute of Technology
Research
Brains have an amazing ability to learn and store information for long periods - in
some cases, a lifetime. A major challenge in neuroscience is to understand how neuronal
networks are sculpted by experience and how proteins/genes contribute to circuit modification.
The goal of our research is to understand information storage, from the molecular
level through in vivo neuronal networks and how these processes go awry in neurological
disorders. My lab utilizes coordinated biochemical, cell biological, electrophysiological
and imaging studies both in vitro and in vivo.
My lab discovered a new and unexpected intercellular communication pathway regulated
by Arc (https://www.sciencedirect.com/science/article/pii/S0092867417315040), a neuronal gene critical for synaptic plasticity, memory, and cognition. Arc contains
a Gag capsid homology domain that has conserved secondary structure with retroviruses
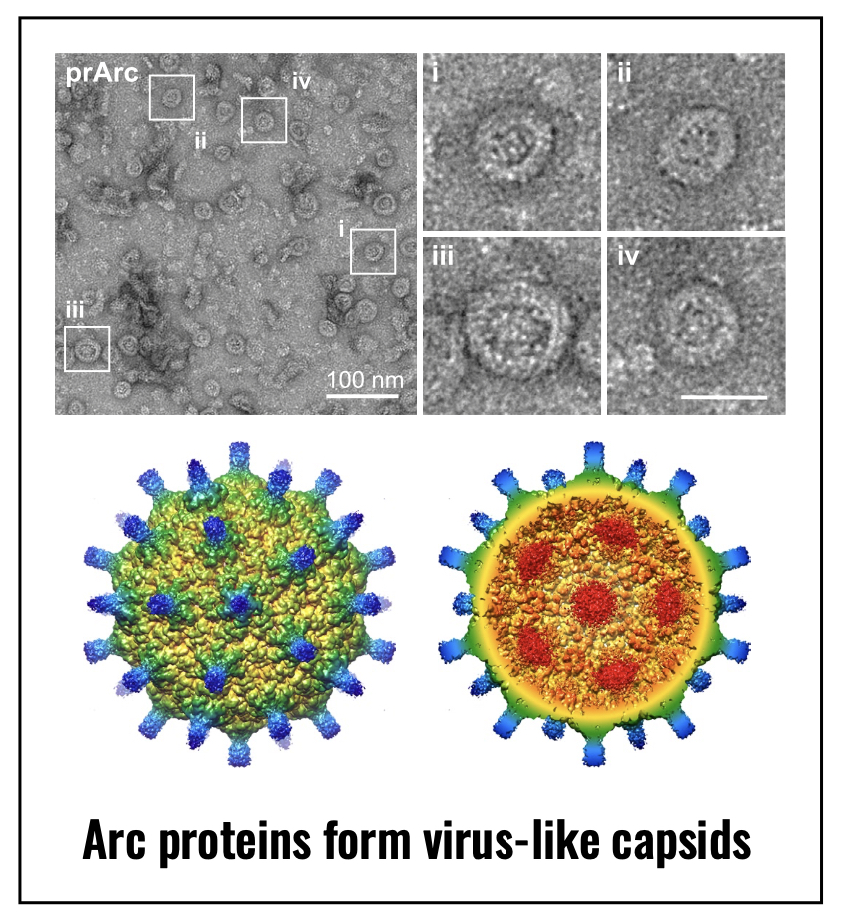
that evolved from a distinct family of retrotransposons. We found that Arc protein
can self-assemble into virus-like capsids that are released from neurons in extracellular
vesicles (EVs), which can transfer RNA and protein cell-to-cell. These findings have
revealed a new form of cell-to-cell communication. A number of Gag-like genes expressed
in various cell types in mammals and humans are predicted to have similar homology,
suggesting that this form of intercellular communication may be present in multiple
cell types and function in many biological systems. In recent work, we found that
the Gag-like paraneoplastic Ma antigen (PNMA) genes encode proteins that are also
capable of capsid formation (https://www.sciencedirect.com/science/article/pii/S0092867424000114). The precise function and signaling pathways mediated by these virus-like processes
promises to reveal new biology that underlies complex brain processes such as memory
and will provide insight into neurological diseases. Ongoing projects in the lab aim
to dissect these new intercellular pathways that intersect diverse fields of biology
that include virology, extracellular communication, evolutionary biology, gene delivery
and neuroscience.
Other projects in the lab include:
1. The role of Arc capsids in memory formation and synaptic plasticity.
2. The normal function of PNMA genes in the nervous system and auto-immune neurological disorders.
3. Harnessing Gag-like genes for gene delivery.
4. Molecular mechanisms of tau pathology in Alzheimer’s Disease.
Other projects in the lab include:
1. The role of Arc capsids in memory formation and synaptic plasticity.
2. The normal function of PNMA genes in the nervous system and auto-immune neurological disorders.
3. Harnessing Gag-like genes for gene delivery.
4. Molecular mechanisms of tau pathology in Alzheimer’s Disease.
References (Selected Publications)
- Xu J#., Erlendsson S., Singh M., Holling G.A., Regier M., Ibiricu I., Einstein J., Hantak M.P., Day G.S., Piquet A.L., Smith T.L., Clardy S.L., Whiteley A.M., Feschotte C., Briggs J. A. G., Shepherd J.D. PNMA2 forms non-enveloped virus-like capsids that trigger paraneoplastic neurological syndrome. Cell. 2024 Feb 15;187(4):831-845.e19. doi: 10.1016/j.cell.2024.01.009. Epub 2024 Jan 31. PMID: 38301645.
- Ravens A., Stacher-Hörndli C.N., Steinwand S., Shepherd J.D., Gregg C. Arc Regulates a “Second-Guessing” Cognitive Bias During Naturalistic Foraging. iScience. 2023 Apr 27;26(5):106761. doi: 10.1016/j.isci.2023.106761.
- Kyrke-Smith M., Volk L.J, Cooke S.F., Bear M.F., Huganir R.L., Shepherd J.D. The immediate early gene Arc is not required for hippocampal long-term potentiation. Journal of Neuroscience, 2021 May 12;41(19):4202-421.
- Hantak M.P., Einstein J., Kearns R.B#.,Shepherd J.D.Intercellular Communication in the Nervous System Goes Viral. Trends in Neurosciences. 2021 Jan 21: S0166-2236(21)00001-1.
- Beagan J.A., Pastuzyn E.D., Fernandez L.R., Titus K.R., Chandrashekar H., Shepherd J.D.*, Phillips-Cremins J.E*. Three-dimensional genome restructuring across timescales of activity-induced neuronal gene expression. Nature Neuroscience. 2020 Jun;23(6):707-717. *Co-corresponding author.
- Jenks K.R., and Shepherd J.D. Experience-dependent development and maintenance of binocular neurons in the mouse visual cortex. Cell Reports, 2020 Feb 11;30(6):1982-1994.e4.
- Erlendsson S., Murado D.R., Cullen H.B, Feschotte C., Shepherd J.D., Briggs J.A.G. Nature Neuroscience, 2020 Feb;23(2):172-175. Structures of virus-like capsids formed by the Drosophila neuronal Arc proteins. Nature Neuroscience, 2020 Feb;23(2):172-175.
- Pastuzyn E.D., Day C.E., Kearns R.B#., Kyrke-Smith M., Taibi A.V., McCormick J., Belnap D.M., Yoder N., Erlendsson S., Morado D.R., Briggs J.A.G., Feschotte C. and Shepherd J.D. The Neuronal Gene Arc Encodes a Repurposed Retrotransposon Gag Protein that Mediates Intercellular RNA Transfer. Cell, 2018 Jan 11;172(1-2):275-288.e18.
#Bioscience student; lab members underlined
*Equal contribution
Links
